Facial weakness: A dark matter detective story

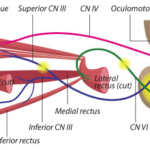
Elizabeth Engle, MD, has devoted her career to finding genetic and developmental causes for disorders of eye, eyelid, and facial movement. From common conditions like strabismus to very rare disorders, these conditions can impact a person’s appearance and impair social communication, making it hard to shift one’s eyes up, down, or sideways or adjust facial expressions.
Each discovery in Engle’s lab offers an answer for families. It’s also a window into the brain — specifically into the development of the nerves and motor neurons that innervate the eye and face muscles.
Now, after the better part of a decade, Engle’s team in the F.M. Kirby Neurobiology Center at Boston Children’s Hospital drove the cracking of a previously impenetrable mystery: the genetic cause of a condition called hereditary congenital facial paresis type 1 (HCFP1).
HCFP1 causes mild facial weakness that makes it hard for babies and children to smile brightly, pucker their lips, suck, and sip from straws. While it’s mild and very rare, the condition has opened the door to a little-known area of genetics: disorders caused by inherited mutations in the genome’s “dark matter.”
A rare category of genetic disease
Most known genetic causes of inherited disease come from variants in genes — the 1 to 2 percent of our genome, also known as the exome, that provides the code for making proteins. But within the rest of the genome — the dark matter — lie important non-coding elements that regulate genes and can influence disease.
“While we only have about 20,000 genes, non-coding regulatory elements provide much additional information, modulating gene expression by turning genes on or off in specific cell types at specific times,” says Engle.
Inherited, single-gene disorders involving non-coding variation remain mostly underexplored, in part because we don’t yet know how to interpret most variation within the vast non-coding space. The new work provides a rare example that could inform future genetic discovery for other disorders.
Examining families’ genetic variants
The team started with nine families from the Engle lab’s database and five families from collaborators in the Netherlands. Linkage mapping in the Dutch families had identified the general area in the genome that seemed tied to HCFP1. The families had had whole-exome sequencing, but since this includes only the coding parts of the genome, it came up empty.
To solve questions like this, one needs a team of people with multiple areas of expertise. We think our work provides an example of how to tease out the genetics and mechanisms of inherited disorders involving noncoding variation.”
Engle’s team used whole genome sequencing, which revealed that five families had a large, duplicated piece of DNA. The start and end points of this piece varied, but in all five families it contained three non-coding regulatory elements. The other nine families had single nucleotide variants (SNVs) — single “letter” changes in the genetic code — all within one of the three regulatory elements.
“That’s a bit of a conundrum, because we typically think of DNA duplications as increasing gene expression, and SNVs as decreasing gene function,” says Engle. “We had to figure out the mechanism by which these duplications and SNVs resulted in the same facial phenotype and determine what gene these regulatory elements were targeting.”
They found that the three regulators are located close to GATA2, a gene known to help specify the production of different cell types in the blood and nervous system. That raised the question: Do the three regulators act on GATA2? Engle’s team suspected they did, and that two enhanced and one silenced GATA2 expression.
Tracking cells’ developmental journeys
That’s where much legwork came in. Using mouse models and tools like single-cell RNA sequencing and CUT&Tag to profile cells at different time points, the team began to understand how variants in DNA regulatory elements lead to facial weakness.
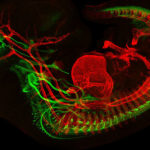
First, they found that the regulatory elements are active during early brain development, in a location in the hindbrain where two cell types are born: facial motor neurons and inner ear efferent neurons (involved in filtering loud sounds and background noise).
“Our hypothesis was that inner ear efferents are born first and are dependent on the enhancers turning on GATA2, and that a day or so later, the silencer shuts down GATA2 and facial motor neurons are made,” says Engle. “We thought that the DNA duplications could increase enhancer activity while the SNVs could reduce silencing activity — both of which would lead to production of more inner ear efferents and fewer facial motor neurons.”
Meticulous experiments showed this to be true. “The combined number of cells was unchanged, but prolonging GATA2 expression increased the inner ear efferents at the expense of the facial motor neurons,” says Alan Tenney, PhD, one of the postdoctoral research fellows in Engle’s lab who led the study.
An explanation for HCFP1
In short, the team showed that the genetic mutations prevent people from making enough motor neurons during development to properly innervate the facial muscles, leaving them relatively weak.
“To solve questions like this, one needs a team of people with multiple complementary areas of expertise,” says Engle. “We think our work provides an example of how to tease out the genetics and mechanisms of inherited disorders involving noncoding variation. The key is what you choose to tackle — which noncoding variants are compelling enough to spend the next five years of your life investigating.”
Tenney, Alessandro Di Gioia, PhD, and Bryn Webb, MD, were co-first authors on the study, published in Nature Genetics.
Learn more about research in the Engle lab and F.M. Kirby Neurobiology Center at Boston Children’s.
Related Posts :
-

Tracking the elusive genes that cause strabismus
Strabismus is a common condition in which the eyes do not align properly, turning inward, outward, upward or downward. Two ...
-

The neurology resident that could
Elizabeth Engle used to wait in peoples' driveways until midnight, hoping to enroll them in her genetic studies of eye-movement ...
-

Two neuroscience rock stars elected to the National Academy of Medicine
Beth Stevens, PhD, and Elizabeth Engle, MD, are the latest Boston Children’s Hospital researchers to be elected to the ...
-

The Beauty of the Brain
Every year, the Harvard Brain Science Initiative sponsors its Beauty of the Brain contest. This year, two Boston Children’s ...