Sturdier spikes may explain SARS-CoV-2 variants’ faster spread
The fast-spreading U.K., South Africa, and Brazil variants are raising concerns and questions about whether current COVID-19 vaccines will protect against them. A structural biology study led by Bing Chen, PhD, at Boston Children’s Hospital now reveals how the D614G mutation — carried by all three variants — makes SARS-CoV-2 spread faster.
Key takeaways:
- The main circulating SARS-CoV-2 variants (U.K., South Africa, Brazil) all carry a genetic change that stabilizes the spike protein, increasing the chances of infection.
- Adding this change to current vaccines could make them better able to elicit protective neutralizing antibodies.
The D614G mutation alters the genetic code for the SARS-CoV-2 spike protein, which is used in most COVID-19 vaccines, by changing a single amino acid “letter.” To see the effects, Chen and colleagues imaged the spikes with cryo-electron microscopy (cryo-EM), which has resolution down to the atomic level. Their findings, reported in Science, suggest a straightforward way to improve vaccines.
The mutation, they found, makes the spikes more stable than those on the original virus. As a result, more functional spikes are available to bind to our cells’ ACE2 receptors, making the virus more infectious.
Preventing spikes’ shape change
To initiate an infection, the spike proteins must bind to the ACE2 receptor. They then dramatically change shape, folding in on themselves. This enables the virus to fuse its membrane with our own cells’ membranes and get inside. However, as Chen and colleagues reported in 2020, the spikes in the original SARS-CoV-2 sometimes prematurely changed shape and fell apart before the virus could bind to cells. While this slowed the virus down, the shape change also made it harder for our immune system to contain it.
“Because the original spike protein would dissociate, it was not good enough to induce a strong neutralizing antibody response,” says Chen.
But when Chen and colleagues imaged the mutant spike protein, they found that the D614G mutation stabilizes the spike by blocking the premature shape change. Interestingly, the mutated spikes bind more weakly to the ACE receptor, but the fact that they’re less apt to fall apart prematurely renders the virus more infectious overall.
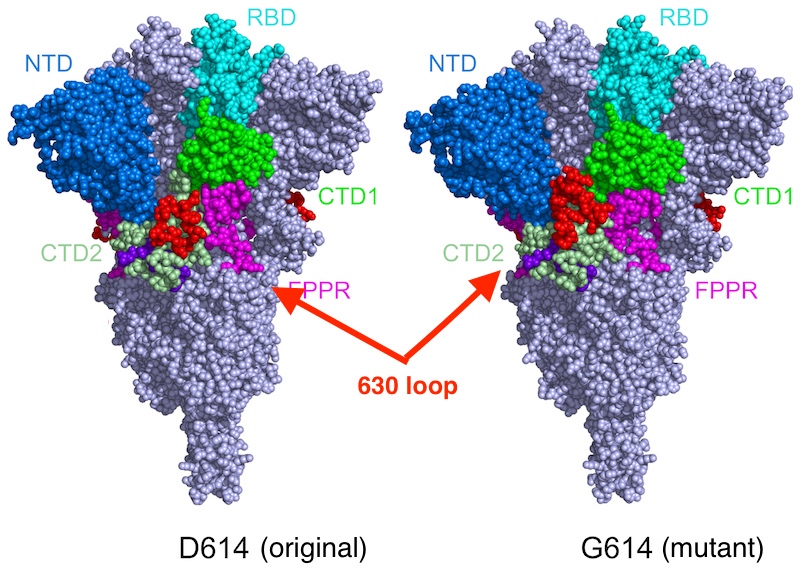
“Say the original virus has 100 spikes,” Chen explains. “Because of the shape instability, you may have just 50 percent of them functional. In the G614 variants, you may have 90 percent that are functional. So even though they don’t bind as well, the chances are greater that you will have infection.”
Future direction: A drug to block coronavirus entry
Chen proposes that redesigned vaccines incorporate the code for this mutant spike protein. He believes the more stable spike shape should make any vaccine based on the spike (including the Moderna, Pfizer, Johnson & Johnson, and AstraZeneca vaccines) more likely to elicit protective neutralizing antibodies.
But Chen also has his sights set on therapeutics. He and his colleagues are further applying structural biology to better understand how SARS-CoV-2 binds to the ACE2 receptor. That could point the way to drugs that would block the virus from gaining entry to our cells.
In January, the team showed that a structurally-engineered “decoy” ACE2 protein binds to SARS-CoV-2 200 times more strongly than the body’s own ACE2. The decoy potently inhibited the virus in cell culture, suggesting it could be an anti-COVID-19 treatment. Chen is now working to advance this research into animal models.
Chen is senior investigator on the paper in Science. Jun Zhang and Yongfei Cai in Boston Children’s Division of Molecular Medicine were co-first authors. The work was funded by the National Institutes of Health, the Massachusetts Consortium on Pathogen Readiness, and Emergent Ventures.
Learn about more COVID-19 research from Boston Children’s.
Related Posts :
-

The hidden burden of solitude: How social withdrawal influences the adolescent brain
Adolescence is a period of social reorientation: a shift from a world centered on parents and family to one shaped ...
-

A new druggable cancer target: RNA-binding proteins on the cell surface
In 2021, research led by Ryan Flynn, MD, PhD, and his mentor, Nobel laureate Carolyn Bertozzi, PhD, opened a new chapter ...
-

No limitations: How Flora found answers for MOG antibody disease
Flora Ringler’s fifth birthday didn’t turn out as she had hoped. She and her family were vacationing in ...
-

What orthopedic trauma surgeons wish more parents knew about lawnmower injuries
Summer is full of delights: lemonade, ice cream, and fresh-cut grass to name a few. Unfortunately, the warmer months can ...