Typing medulloblastoma: From RNA to proteomics and phospho-proteomics
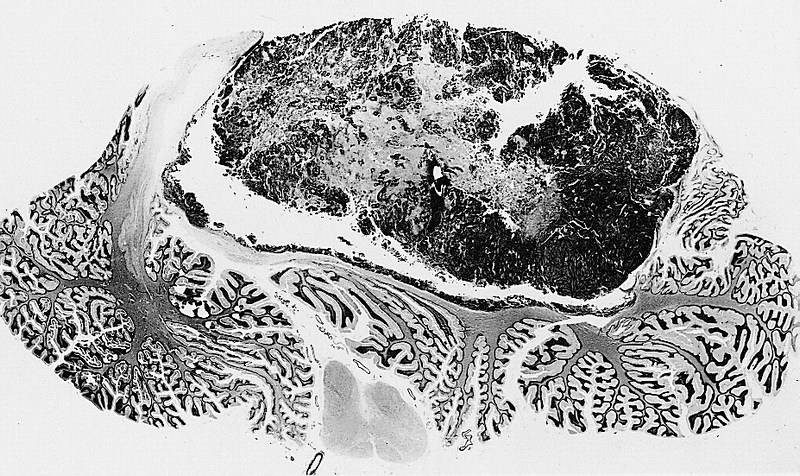
Medulloblastoma is one of the most common pediatric brain tumors, accounting for nearly 10 percent of cases. It occurs in the cerebellum, a complex part of the brain that controls balance, coordination and motor function and regulates verbal expression and emotional modulation. While overall survival rates are high, current therapies can be toxic and cause secondary cancers. Developing alternative therapeutics is a priority for the field.
As early as the 1990s, the lab of Scott Pomeroy, MD, PhD, neurologist-in-chief at Boston Children’s Hospital, discovered molecules in medulloblastoma tumors that could predict response to therapies. In 2010, Pomeroy and colleagues uncovered four distinct molecular subtypes of medulloblastoma.
The World Health Organization updated the brain tumor classification scheme in 2016 to include these molecular and genetic features. In the new scheme, tumor subtypes with a good molecular prognosis receive less radiation and chemotherapy. But the creation of targeted therapeutics has remained a challenge, since some of the genetic pathways implicated in these subtypes are found in non-cancerous cells.
Searching for biological significance
A new study, published yesterday in Cancer Cell, turned to proteomics, the large-scale study of proteins in cells. It also looked at the enzymes involved in phosphorylation, the addition of a phosphate group to a molecule and a major mechanism by which proteins are regulated.
Previous tumor typing looked at RNA, which is made from DNA and which the cell “translates” to make proteins. But as the new study showed, there is not necessarily a one-to-one correlation between RNA and the protein it translates.
“We hypothesized that the relationship of protein quantity and its RNA template is not random,” says Pomeroy, a senior author on the paper.
Leveraging proteomics, Pomeroy and colleagues showed that the protein-versus-RNA ratio varies for each medulloblastoma subtype. They also discovered differences in which proteins are actually present from tumor to tumor and how the proteins interact with other molecules in the cell once they’re made.
The results, extending more than 20 years of findings from Pomeroy’s lab, offer a glimpse at potential therapeutic targets in medulloblastoma.
Zeroing in on modified proteins
Pomeroy’s team worked with groups led by Ernest Fraenkel, PhD, professor of biological engineering in MIT’s Department of Biological Engineering, Steven Carr, PhD, senior director of proteomics at the Broad Institute of MIT and Harvard, and Jill Mesirov, PhD, of the Broad Institute and the Moores Cancer Center of UC San Diego to profile 45 medulloblastoma samples. They found extensive “remodeling” of the proteome after translation, even in tumor subtypes with similar patterns of RNA expression.
Since proteins direct biological activities in tumor cells, proteomic and phospho-proteomic analyses are effective ways of identifying potential therapeutic targets.
They discovered that MYC proteins, which are known to drive the growth of cancers, were chemically modified in a subset of the group 3 tumors that had especially poor outcomes. By tracing these protein modifications to their source, the phospho-proteomic analysis pointed to a kinase called PRKDC that plays a role in DNA stability. Inhibiting PRKDC with an existing chemical in these tumor cells sensitized the cells to radiation.
“Although the PRKDC inhibitor has yet to be tested in humans, this could potentially be a viable approach to treating a medulloblastoma subtype with historically poor outcomes,” Pomeroy says.
“Since proteins direct biological activities in tumor cells, global proteomic and phospho-proteomic analyses are effective ways of identifying potential therapeutic targets,” he adds. “We are hopeful that the next phase of our research will bring us even closer to developing viable treatments.”
More information in this press release from the Broad Institute. See the paper for a full list of authors and funders.
Related Posts :
-

Mapping ‘neighborhoods’ in aggressive childhood brain tumors
The third-most common kind of childhood brain tumor, supratentorial ependymoma, is an aggressive cancer that often returns after surgery and ...
-

Gene therapy for hearing loss: Tag-teaming from the lab to the clinic
Two-year-old Miles is one of the first patients with hereditary hearing loss to receive gene therapy at Boston Children’s ...
-

‘Challenge accepted’: Sophia takes on a brain tumor
In 2023, Sophia Mordini landed the role of a lifetime. A competitive dancer, the 12-year-old would play Clara in her company’...
-

A true hero’s journey: How a team approach helped Wolfie overcome pancreatitis
Wolfgang, affectionately known as “Wolfie,” is a bright and energetic 7-year-old with a quick wit and a love for making ...