Stem cell medicine gets a “roadmap” and a quality assurance tool
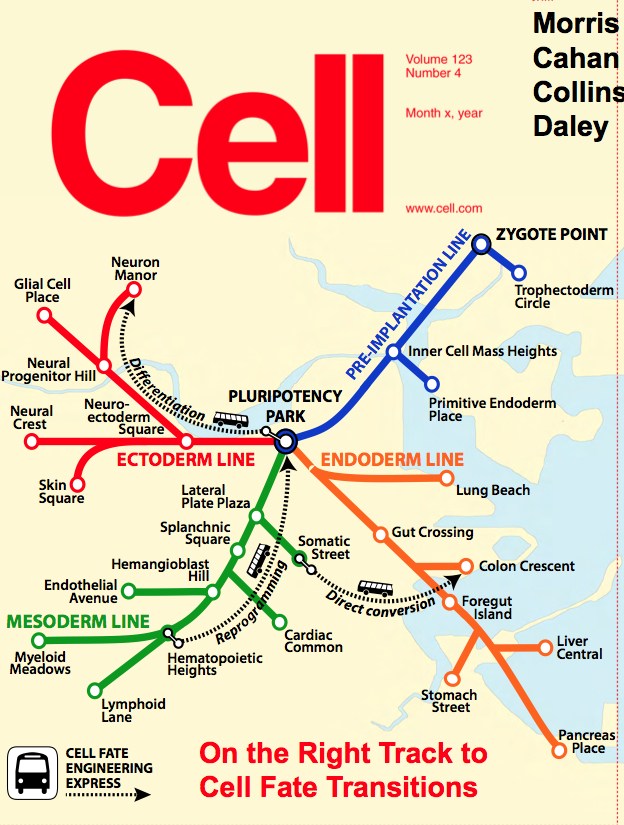
If you’ve lost your way on the Boston subway, you need only consult a map to find the best route to your destination. Now stem cell engineers have a similar map to guide the making of cells and tissues for disease modeling, drug testing and regenerative medicine. It’s a computer algorithm known as CellNet.
As in this map on the cover of Cell, a cell has many possible destinations or “fates,” and can arrive at them through three main stem cell engineering methods:
• reprogramming (dialing a specialized cell, such as a skin cell, back to a stem-like state with full tissue-making potential)
• differentiation (pushing a stem cell to become a particular cell type, such as a blood cell)
• direct conversion (changing one kind of specialized cell to another kind)
Freely available on the Internet, CellNet provides clues to which methods of cellular engineering are most effective—and acts as a much-needed quality control tool.
“To date, there has been no systematic means to determine how closely cells made in a petri dish approximate natural tissues in the body,” says George Q. Daley, MD, PhD, director of the Stem Cell Transplantation Program at Boston Children’s Hospital, senior investigator on two studies published by Cell last week.
CellNet adds that analytical rigor and even suggests ways to make the cells better. As shown below, the algorithm’s “inputs” are engineered cells made through the different methods. The “outputs” are comparisons of these cells’ gene regulatory networks (which genes are turned on or off) to those of the real-life cells or tissues they’re meant to emulate. At far right, the algorithm flags potential genetic on/off switches that a scientist could target to improve upon his or her cells, then ranks them in order of priority.

“CellNet will also be a powerful tool to advance synthetic biology—to engineer cells for specific medical applications,” says James Collins, PhD, of the Wyss Institute for Biologically Inspired Engineering and Boston University, and co-senior author on the first study, which used CellNet to assess cells created in 56 published studies.
The second study delved into a recurring question in stem cell biology: Is it feasible to directly convert one specialized cell type to another, skipping the laborious process of making a stem cell?
“Previously, most attempts to directly convert one specialized cell type to another have depended on a trial-and-error approach,” notes Patrick Cahan, PhD, a Daley lab member and principal architect of CellNet.
The new research confirms that most directly converted cells retain some “memory” of their cell of origin. That makes them less than ideal for certain uses—but surprisingly better for others.
For example, when the researchers tried to turn skin cells into liver cells, their effort failed—but unexpectedly yielded cells closely resembling intestinal cells.
“The converted skin cells were able to engraft into mice with inflammatory bowel disease—Crohn’s or ulcerative colitis,” says co-first author Samantha Morris, PhD, a researcher in the Daley lab. “After a short time, the cells became highly similar to native colon cells and assisted healing of the damaged tissue.”
Related Posts :
-

A case for Kennedy — and for rapid genomic testing in every NICU
Kennedy was born in August 2025 after what her parents, John and Diana, describe as an uneventful pregnancy. Soon after delivery, ...
-

The journey to a treatment for hereditary spastic paraplegia
In 2016, Darius Ebrahimi-Fakhari, MD, PhD, then a neurology fellow at Boston Children’s Hospital, met two little girls with spasticity ...
-

New research paves the way to a better understanding of telomeres
Much the way the caps on the ends of a shoelace prevent it from fraying, telomeres — regions of repetitive DNA ...
-

AI-designed proteins open doors to new immunotherapies
Artificial intelligence (AI) is increasingly helping drive advances in science and medicine — including cellular signaling. In a recent study, published ...





