Families and data scientists build insights on Phelan-McDermid syndrome

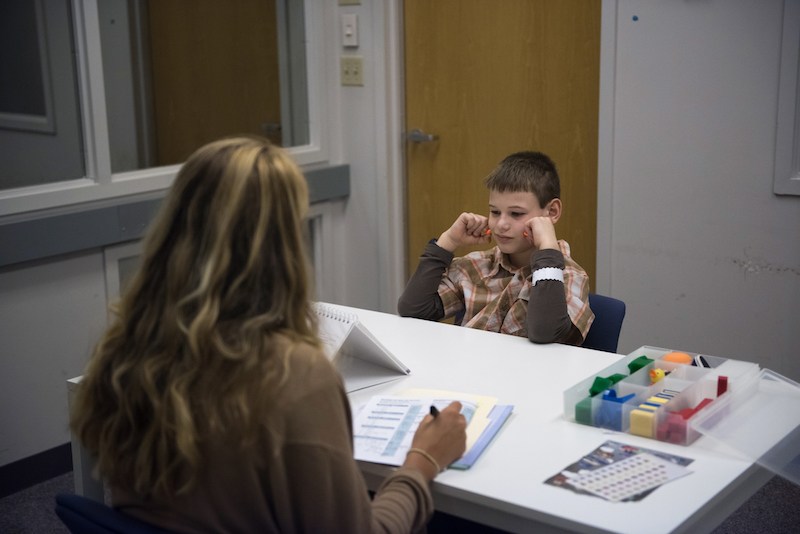
This is the third year that Jacob Works has made the trip down to Boston Children’s Hospital from Maine. With research assistant Haley Medeiros, he looks at pictures, answers questions, manipulates blocks and mimes actions like knocking on a door. His father, Travis, and another research assistant look on through a window.
“At first, we had to practically bribe him with an iPad with every task,” Travis says. “This year he’s more excited, because he understands more and is more confident and able to share more.”
Jacob, 11, was diagnosed in 2011 with Phelan-McDermid Syndrome, a rare genetic condition that typically causes children to be born “floppy,” with low muscle tone, and to have little or no speech, developmental delay and, often, autism-like behaviors. At the time, Jacob was one of about 800 known cases. But through chromosomal microarray testing, introduced in just the past decade for children with autism symptoms, more cases are being picked up.
Phelan-McDermid, also known as PMS, 22q13 deletion syndrome or 22q13.3 deletion syndrome, is caused by loss of a piece of DNA in chromosome 22, or a “misspelling” in the SHANK3 gene. (SHANK3 is sometimes also deleted as part of the chromosome deletion).

“There’s a great variability in presentation,” says Siddharth Srivastava, MD, who sees children with Phelan-McDermid Syndrome in Boston Children’s weekly developmental neurogenetics clinic. “Some kids are nonverbal, with very limited language communication, and some are less affected.”
Jacob is at the mild end of the spectrum; low muscle tone has altered his gait slightly and he was late to begin speaking, but he is quite verbal today. He’s one of 100 children in a long-term “natural history” study of the NIH-funded Developmental Synaptopathies Consortium. Led by Boston Children’s Hospital, the study’s goal is to map children’s genetic information to their symptoms and evaluation findings over time. Mustafa Sahin, MD, PhD, at Boston Children’s is lead principal investigator.
“There’s not a whole lot of information out there,” says Works. “Anything we can do to contribute information toward understanding the disorder, we want to do.”
A “Google Maps” for Phelan-McDermid?
In Google Maps, you can zoom in and see traffic jams between you and your destination, locate nearby restaurants and find out their hours. Parents Geraldine Bliss and Megan O’Boyle from the Phelan-McDermid Syndrome Foundation (PMSF), together with data scientist Paul Avillach, MD, PhD, have accomplished an analogous feat for Phelan-McDermid Syndrome.
In 2011, the PMSF created the Phelan-McDermid International Registry. It allows families to input their children’s medical information, prompted by 300 questions, and freely retrieve it. Of the 1,800 patients diagnosed with the syndrome to date, more than 1,100 are enrolled.
But Bliss and O’Boyle wanted to make even more data available to researchers, to help them better understand Phelan-McDermid and develop treatments.
A blue button
Around this time, the Patient-Centered Outcomes Research Institute (PCORI) was funding “blue button” initiatives to help patients access and use their data. But to apply for a grant, the Foundation needed an academic partner.
Searching “blue button” and “autism” online, Bliss found Isaac Kohane, MD, PhD, head of the Computational Health Informatics Program (CHIP) at Boston Children’s and chair of the Department of Biomedical Informatics (DBMI) at Harvard Medical School. Kohane steered Bliss and O’Boyle to Avillach, who was looking for a good “big data” challenge.
Avillach left his tenured position in France to join the faculty at Boston Children’s Hospital and HMS’s Department of Biomedical Informatics. He and the PMSF got a $1 million PCORI grant in 2013, and the Phelan-McDermid Syndrome Data Network, or PMS_DN, was born. O’Boyle is principal investigator.
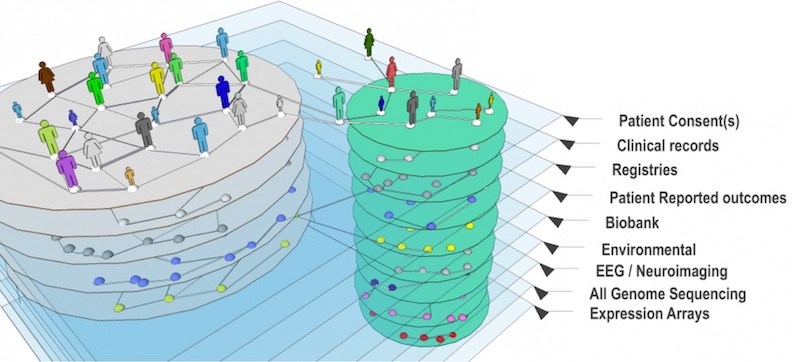
Centralizing Phelan-McDermid patient data
The PMS_DN is multi-layered. In addition to patient registry data, it pulls in medical records, diagnostic codes, doctors’ notes and data from genetic reports. More than 700 patients have joined.
To gather their clinical records and notes, Avillach and the Foundation tapped a service called CareSync. Families sign a release form for each provider, and CareSync does all the legwork.
That was a crucial step. “The data you can get from a healthcare system’s patient portal — things like vaccination records — aren’t enough data for our families,” O’Boyle says. “Parents shouldn’t have to guess when their child walked, talked and had their first seizure.”
O’Boyle recalls having to call six separate hospitals, fill out paperwork and pay up to $1.43 per page to get medical records for her daughter Shannon, now 17 — then carrying this sheaf of records to every doctor visit.
Now Shannon’s data — and everyone else’s — are in one central, accessible place.
“Usually in big data, we have a lot of patients,” Avillach says. “In this case, we have only 750 patients, but more than 20 million clinical data points.”
Deep phenotyping
Avillach has layered on tools to organize and integrate the data, including i2b2/tranSMART, a “knowledge management” platform; RESTful API, which helps extract patient data from medical records; PIC-SURE and Jupyter Notebook.
Key is a natural-language-processing algorithm called CTAKES, developed at Boston Children’s and the Mayo Clinic. It captures clinician-written notes and referral letters and converts them into standardized data. For example, “biliary calculus” and “gallstones” are classified as the same thing.
The PMS_DN can be queried with different levels of access. Any investigator working on autism and/or intellectual disability can request Level 1 access, where questions can be asked in the aggregate, like “If patients have seizures, what other complications do they have?” These general queries can help shape more focused research questions. Investigators wanting patient-level data can request it with a research project authorized by an institutional review board (IRB).
The list of potential questions is endless: What is the average age of diagnosis? What is the age of onset of different symptoms? How commonly are skills lost? How often are they regained? How do symptoms differ between children with different chromosome 22 deletions, versus those with mutations in SHANK3 alone?
“We built a phenomenal tool and we’ll get it into the hands of the research community and they can start doing queries,” says O’Boyle. “Hopefully, we’ll be involved in drug development as well.”
Related Posts :
-

Gene therapy for hearing loss: Tag-teaming from the lab to the clinic
Two-year-old Miles is one of the first patients with hereditary hearing loss to receive gene therapy at Boston Children’s ...
-

Promising advances in fetal therapy for vein of Galen malformation
In 2024, Megan Ingram* of California and her husband were preparing for the birth of their third child when a 34-week ...
-

A case for Kennedy — and for rapid genomic testing in every NICU
Kennedy was born in August 2025 after what her parents, John and Diana, describe as an uneventful pregnancy. Soon after delivery, ...
-

The hidden burden of solitude: How social withdrawal influences the adolescent brain
Adolescence is a period of social reorientation: a shift from a world centered on parents and family to one shaped ...





